Our Research
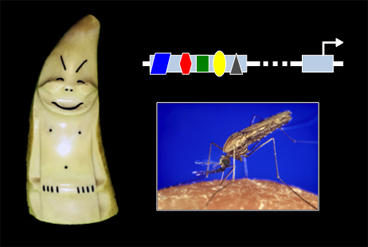
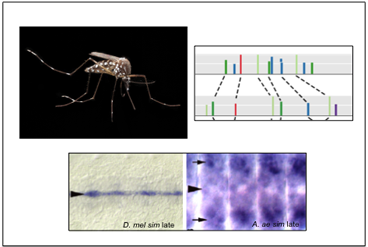
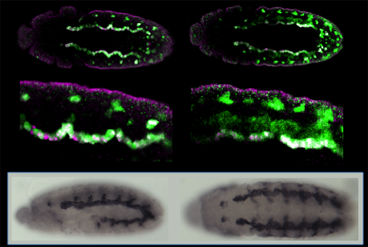
While all cells in an organism share the same genome, how that genome is used—which genes are turned "on" or "off" at any given time, for instance—needs to be tightly regulated. Understanding this regulation, and how it evolved, is fundamental to understanding how cell fates are determined during development, and for acquiring insight into normal variation between individuals, birth defects, chronic diseases, and evolution. We use a combination of molecular, genetic, and computational approaches to study the regulatory genomics of the fruitfly Drosophila melanogaster and other insect species of both medical (e.g., mosquitoes) and agricultural importance.
The projects in our laboratory respectively investigate enhancers, promoters, and intercellular signals—the three basic building blocks of developmental regulatory networks. As we progress further into understanding each of these components and develop new tools and methods for their investigation, we move closer to being able to piece all three together into a coherent "systems level" picture of how cell fates are determined as an organism develops, and of the consequences when development goes awry.